CRISPR-U™ Gene Knockout Cell Line
Strategy
MAP4 Gene Knockout Strategy
CRISPR-U™ technology (CRISPR based), developed by Ubigene, is more efficient than general
CRISPR/Cas9 technology in double-strand breaking and homologous recombination. With CRISPR-U™, Ubigene has
successfully edited over 3000 genes on more than 100 types of cell lines.
Objective
To create a Human MAP4 Knockout
model in cell line by
CRISPR-U™-mediated
genome
engineering.
Target gene info 
| Official symbol | MAP4 |
| Gene id | 4134 |
| Organism | Homo sapiens |
| Official full symbol | microtubule associated protein 4 |
| Gene type | protein-coding |
| Summary | The protein encoded by this gene is a major non-neuronal microtubule-associated protein. This protein contains a domain similar to the microtubule-binding domains of neuronal microtubule-associated protein (MAP2) and microtubule-associated protein tau (MAPT/TAU). This protein promotes microtubule assembly, and has been shown to counteract destabilization of interphase microtubule catastrophe promotion. Cyclin B was found to interact with this protein, which targets cell division cycle 2 (CDC2) kinase to microtubules. The phosphorylation of this protein affects microtubule properties and cell cycle progression. Multiple transcript variants encoding different isoforms have been found for this gene. |
| Genomic regions | Chromosome 3 |
Strategy Summary
This gene has 12 protein coding transcripts:
| Name | Transcript ID | bp | Protein | Biotype | CCDS | UniProt Match | RefSeq Match | Flags |
| MAP4-204 | ENST00000395734.7 | 5590 | 1135aa | Protein coding | CCDS46818 | P27816-6 | - | TSL:2, GENCODE basic, APPRIS ALT2, |
| MAP4-202 | ENST00000360240.10 | 5142 | 1152aa | Protein coding | CCDS33750 | P27816-1 | - | TSL:1, GENCODE basic, APPRIS ALT2, |
| MAP4-209 | ENST00000434267.5 | 2772 | 99aa | Protein coding | CCDS46821 | P27816-7 | - | TSL:2, GENCODE basic, |
| MAP4-210 | ENST00000439356.2 | 2743 | 99aa | Protein coding | CCDS46821 | P27816-7 | - | TSL:1, GENCODE basic, |
| MAP4-207 | ENST00000426837.6 | 8920 | 2297aa | Protein coding | - | E7EVA0 | - | TSL:5, GENCODE basic, APPRIS P3, |
| MAP4-208 | ENST00000429422.5 | 3724 | 493aa | Protein coding | - | H7C4C5 | - | CDS 5' incomplete, TSL:1, |
| MAP4-205 | ENST00000420772.6 | 3478 | 828aa | Protein coding | - | F8W9U4 | - | TSL:5, GENCODE basic, APPRIS ALT2, |
| MAP4-203 | ENST00000383736.3 | 2282 | 760aa | Protein coding | - | B5MEG9 | - | CDS 5' and 3' incomplete, TSL:5, |
| MAP4-201 | ENST00000335271.9 | 1564 | 463aa | Protein coding | - | H0Y2V1 | - | CDS 5' incomplete, TSL:2, |
| MAP4-206 | ENST00000423088.5 | 899 | 283aa | Protein coding | - | H7C456 | - | CDS 3' incomplete, TSL:3, |
| MAP4-216 | ENST00000633710.1 | 570 | 140aa | Protein coding | - | A0A0J9YW37 | - | CDS 3' incomplete, TSL:5, |
| MAP4-212 | ENST00000468075.2 | 487 | 76aa | Protein coding | - | A0A0J9YVV8 | - | CDS 3' incomplete, TSL:5, |
| MAP4-215 | ENST00000497735.1 | 2156 | No protein | Retained intron | - | - | - | TSL:2, |
| MAP4-211 | ENST00000462206.1 | 572 | No protein | Retained intron | - | - | - | TSL:4, |
| MAP4-214 | ENST00000482752.1 | 386 | No protein | Retained intron | - | - | - | TSL:3, |
| MAP4-213 | ENST00000477765.1 | 284 | No protein | Retained intron | - | - | - | TSL:3, |




Strategy
Click to get 
Red Cotton™ Assessment
Project Difficulty Level unknown
| Target Gene | MAP4 |
| This KO Strategy | loading |
| Red Cotton™ Notes | Gene MAP4 had been KO in hela cell line. |
Aforementioned information comes from Ubigene database. Different origin of cell lines may have
different condition. Ubigene reserved all the right for final explanation.
Special deals for this gene:
$49
Single gRNA plasmid off-shelf
$599
Single gRNA lentivirus



Work flow
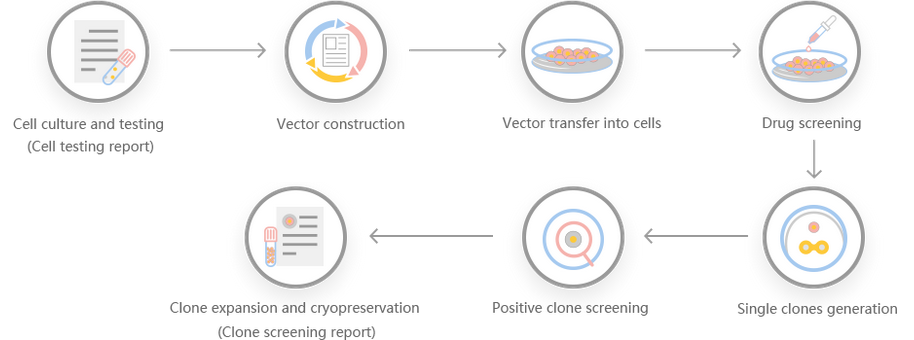





 Comment:
Comment: