CRISPR-U™ Gene Knockout Cell Line
Strategy
ARHGDIA Gene Knockout Strategy
CRISPR-U™ technology (CRISPR based), developed by Ubigene, is more efficient than general
CRISPR/Cas9 technology in double-strand breaking and homologous recombination. With CRISPR-U™, Ubigene has
successfully edited over 3000 genes on more than 100 types of cell lines.
Objective
To create a Human ARHGDIA Knockout
model in cell line by
CRISPR-U™-mediated
genome
engineering.
Target gene info 
| Official symbol | ARHGDIA |
| Gene id | 396 |
| Organism | Homo sapiens |
| Official full symbol | Rho GDP dissociation inhibitor alpha |
| Gene type | protein-coding |
| Also known as | GDIA1, HEL-S-47e, NPHS8, RHOGDI, RHOGDI-1 |
| Summary | This gene encodes a protein that plays a key role in the regulation of signaling through Rho GTPases. The encoded protein inhibits the disassociation of Rho family members from GDP (guanine diphosphate), thereby maintaining these factors in an inactive state. Activity of this protein is important in a variety of cellular processes, and expression of this gene may be altered in tumors. Mutations in this gene have been found in individuals with nephrotic syndrome, type 8. Alternate splicing results in multiple transcript variants. |
| Genomic regions | Chromosome 17 |
Strategy Summary
This gene has 8 protein coding transcripts:
| Name | Transcript ID | bp | Protein | Biotype | CCDS | UniProt Match | RefSeq Match | Flags |
| ARHGDIA-201 | ENST00000269321.12 | 1837 | 204aa | Protein coding | CCDS11788 | P52565-1 | NM_004309.6 | TSL:1, GENCODE basic, APPRIS P1, MANE Select v0.92, |
| ARHGDIA-202 | ENST00000400721.8 | 1568 | 160aa | Protein coding | CCDS58609 | P52565-2 | - | TSL:2, GENCODE basic, |
| ARHGDIA-203 | ENST00000541078.6 | 1517 | 204aa | Protein coding | CCDS11788 | P52565-1 | - | TSL:3, GENCODE basic, APPRIS P1, |
| ARHGDIA-207 | ENST00000580685.5 | 1201 | 204aa | Protein coding | CCDS11788 | P52565-1 | - | TSL:1, GENCODE basic, APPRIS P1, |
| ARHGDIA-217 | ENST00000584461.5 | 886 | 235aa | Protein coding | CCDS77133 | J3QQX2 | - | TSL:2, GENCODE basic, |
| ARHGDIA-215 | ENST00000583868.5 | 815 | 249aa | Protein coding | - | J3KRY1 | - | CDS 3' incomplete, TSL:3, |
| ARHGDIA-208 | ENST00000581876.5 | 709 | 129aa | Protein coding | - | J3KRE2 | - | TSL:3, GENCODE basic, |
| ARHGDIA-205 | ENST00000579121.5 | 637 | 193aa | Protein coding | - | J3KTF8 | - | CDS 3' incomplete, TSL:3, |
| ARHGDIA-206 | ENST00000580033.5 | 547 | 92aa | Nonsense mediated decay | - | J3KS60 | - | TSL:4, |
| ARHGDIA-204 | ENST00000578351.1 | 421 | 92aa | Nonsense mediated decay | - | J3KS60 | - | TSL:4, |
| ARHGDIA-210 | ENST00000582520.1 | 245 | No protein | Processed transcript | - | - | - | TSL:3, |
| ARHGDIA-211 | ENST00000582984.5 | 823 | No protein | Retained intron | - | - | - | TSL:3, |
| ARHGDIA-209 | ENST00000582309.1 | 628 | No protein | Retained intron | - | - | - | TSL:2, |
| ARHGDIA-216 | ENST00000584397.1 | 622 | No protein | Retained intron | - | - | - | TSL:2, |
| ARHGDIA-212 | ENST00000583111.5 | 574 | No protein | Retained intron | - | - | - | TSL:2, |
| ARHGDIA-213 | ENST00000583499.1 | 560 | No protein | Retained intron | - | - | - | TSL:2, |
| ARHGDIA-214 | ENST00000583791.1 | 351 | No protein | Retained intron | - | - | - | TSL:2, |




Red Cotton™ Assessment
Project Difficulty Level unknown
| Target Gene | ARHGDIA |
| This KO Strategy | loading |
| Red Cotton™ Notes | Gene ARHGDIA had been KO in hek293t cell line. |
Aforementioned information comes from Ubigene database. Different origin of cell lines may have
different condition. Ubigene reserved all the right for final explanation.
Special deals for this gene:
$49
Single gRNA plasmid off-shelf
$599
Single gRNA lentivirus



Work flow
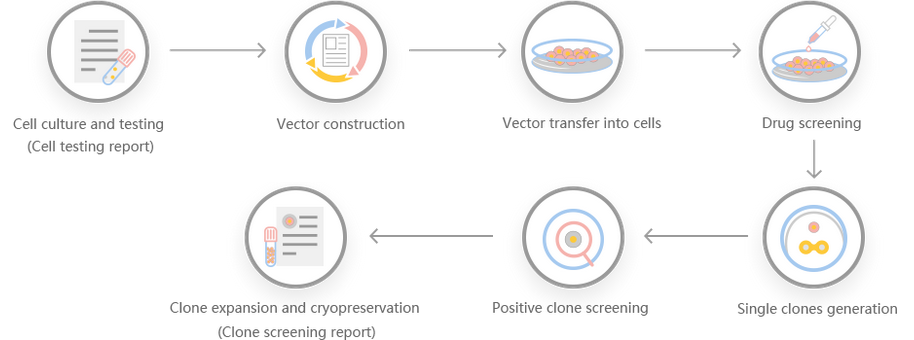






 Comment:
Comment: