CRISPR-U™ Gene Knockout Cell Line
Strategy
GPSM2 Gene Knockout Strategy
CRISPR-U™ technology (CRISPR based), developed by Ubigene, is more efficient than general
CRISPR/Cas9 technology in double-strand breaking and homologous recombination. With CRISPR-U™, Ubigene has
successfully edited over 3000 genes on more than 100 types of cell lines.
Objective
To create a Human GPSM2 Knockout
model in cell line by
CRISPR-U™-mediated
genome
engineering.
Target gene info 
| Official symbol | GPSM2 |
| Gene id | 29899 |
| Organism | Homo sapiens |
| Official full symbol | G protein signaling modulator 2 |
| Gene type | protein-coding |
| Also known as | CMCS, DFNB82, LGN, PINS |
| Summary | The protein encoded by this gene belongs to a family of proteins that modulate activation of G proteins, which transduce extracellular signals received by cell surface receptors into integrated cellular responses. The N-terminal half of this protein contains 10 copies of leu-gly-asn (LGN) repeat, and the C-terminal half contains 4 GoLoco motifs, which are involved in guanine nucleotide exchange. This protein may play a role in neuroblast division and in the development of normal hearing. Mutations in this gene are associated with autosomal recessive nonsyndromic deafness (DFNB82). Alternative splicing results in multiple transcript variants. |
| Genomic regions | Chromosome 1 |
Strategy Summary
This gene has 16 protein coding transcripts:
| Name | Transcript ID | bp | Protein | Biotype | CCDS | UniProt Match | RefSeq Match | Flags |
| GPSM2-202 | ENST00000406462.6 | 7433 | 684aa | Protein coding | CCDS792 | A0A024R0F8 P81274 | - | TSL:5, GENCODE basic, APPRIS P1, |
| GPSM2-201 | ENST00000264126.9 | 7152 | 684aa | Protein coding | CCDS792 | A0A024R0F8 P81274 | NM_013296.5 | TSL:1, GENCODE basic, APPRIS P1, MANE Select v0.92, |
| GPSM2-207 | ENST00000642355.1 | 5609 | 684aa | Protein coding | CCDS792 | A0A024R0F8 P81274 | - | GENCODE basic, APPRIS P1, |
| GPSM2-206 | ENST00000446797.2 | 3194 | 684aa | Protein coding | CCDS792 | A0A024R0F8 B0QZC9 | - | TSL:4, GENCODE basic, APPRIS P1, |
| GPSM2-211 | ENST00000645164.2 | 2806 | 684aa | Protein coding | CCDS792 | A0A024R0F8 A0A2R8Y896 | - | GENCODE basic, APPRIS P1, |
| GPSM2-223 | ENST00000676184.1 | 2741 | 684aa | Protein coding | CCDS792 | A0A024R0F8 | - | GENCODE basic, APPRIS P1, |
| GPSM2-217 | ENST00000675086.1 | 2991 | 625aa | Protein coding | - | - | - | GENCODE basic, |
| GPSM2-218 | ENST00000675087.1 | 2812 | 701aa | Protein coding | - | - | - | GENCODE basic, |
| GPSM2-216 | ENST00000674914.1 | 2700 | 701aa | Protein coding | - | - | - | GENCODE basic, |
| GPSM2-214 | ENST00000674700.1 | 2447 | 519aa | Protein coding | - | - | - | GENCODE basic, |
| GPSM2-205 | ENST00000441735.2 | 2378 | 627aa | Protein coding | - | H0Y4A4 | - | TSL:2, GENCODE basic, |
| GPSM2-209 | ENST00000643643.1 | 1520 | 311aa | Protein coding | - | A0A2R8Y6E3 | - | CDS 5' incomplete, |
| GPSM2-203 | ENST00000435475.5 | 992 | 75aa | Protein coding | - | B0QZD0 | - | CDS 3' incomplete, TSL:3, |
| GPSM2-204 | ENST00000435987.5 | 904 | 213aa | Protein coding | - | Q5T1N9 | - | CDS 3' incomplete, TSL:3, |
| GPSM2-208 | ENST00000643094.1 | 799 | 93aa | Protein coding | - | A0A2R8Y673 | - | CDS 3' incomplete, |
| GPSM2-212 | ENST00000645255.1 | 685 | 228aa | Protein coding | - | A0A2R8YCX1 | - | CDS 5' and 3' incomplete, |
| GPSM2-215 | ENST00000674731.1 | 3841 | 384aa | Nonsense mediated decay | - | - | - | - |
| GPSM2-224 | ENST00000676404.1 | 2472 | 465aa | Nonsense mediated decay | - | - | - | - |
| GPSM2-219 | ENST00000675617.1 | 2386 | No protein | Processed transcript | - | - | - | - |
| GPSM2-213 | ENST00000674663.1 | 1549 | No protein | Processed transcript | - | - | - | - |
| GPSM2-210 | ENST00000643921.1 | 563 | No protein | Processed transcript | - | - | - | - |
| GPSM2-221 | ENST00000675776.1 | 8071 | No protein | Retained intron | - | - | - | - |
| GPSM2-222 | ENST00000675829.1 | 3284 | No protein | Retained intron | - | - | - | - |
| GPSM2-220 | ENST00000675740.1 | 2107 | No protein | Retained intron | - | - | - | - |




Strategy
Click to get 
Red Cotton™ Assessment
Project Difficulty Level unknown
| Target Gene | GPSM2 |
| This KO Strategy | loading |
| Red Cotton™ Notes | Gene GPSM2 had been KO in hela cell line. |
Aforementioned information comes from Ubigene database. Different origin of cell lines may have
different condition. Ubigene reserved all the right for final explanation.
Special deals for this gene:
$49
Single gRNA plasmid off-shelf
$599
Single gRNA lentivirus



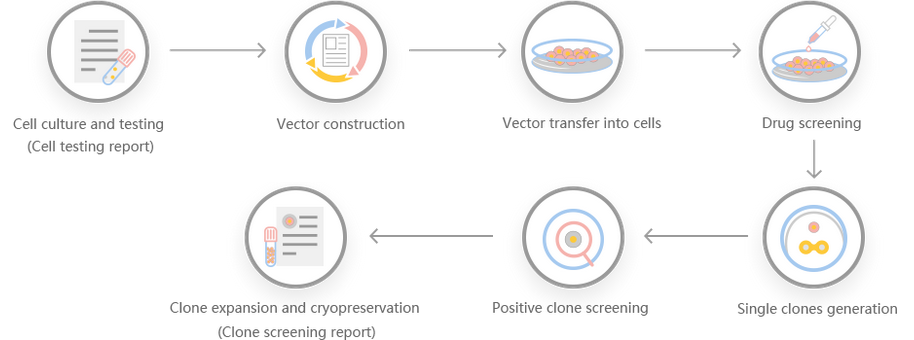
Work flow





 Comment:
Comment: