CRISPR-U™ Gene Knockout Cell Line
Strategy
PEG10 Gene Knockout Strategy
CRISPR-U™ technology (CRISPR based), developed by Ubigene, is more efficient than general
CRISPR/Cas9 technology in double-strand breaking and homologous recombination. With CRISPR-U™, Ubigene has
successfully edited over 3000 genes on more than 100 types of cell lines.
Objective
To create a Human PEG10 Knockout
model in cell line by
CRISPR-U™-mediated
genome
engineering.
Target gene info 
| Official symbol | PEG10 |
| Gene id | 23089 |
| Organism | Homo sapiens |
| Official full symbol | paternally expressed 10 |
| Gene type | protein-coding |
| Also known as | EDR, HB-1, MEF3L, Mar2, Mart2, RGAG3, RTL2, SIRH1 |
| Summary | This is a paternally expressed imprinted gene that is thought to have been derived from the Ty3/Gypsy family of retrotransposons. It contains two overlapping open reading frames, RF1 and RF2, and expresses two proteins: a shorter, gag-like protein (with a CCHC-type zinc finger domain) from RF1; and a longer, gag/pol-like fusion protein (with an additional aspartic protease motif) from RF1/RF2 by -1 translational frameshifting (-1 FS). While -1 FS has been observed in RNA viruses and transposons in both prokaryotes and eukaryotes, this gene represents the first example of -1 FS in a eukaryotic cellular gene. This gene is highly conserved across mammalian species and retains the heptanucleotide (GGGAAAC) and pseudoknot elements required for -1 FS. It is expressed in adult and embryonic tissues (most notably in placenta) and reported to have a role in cell proliferation, differentiation and apoptosis. Overexpression of this gene has been associated with several malignancies, such as hepatocellular carcinoma and B-cell lymphocytic leukemia. Knockout mice lacking this gene showed early embryonic lethality with placental defects, indicating the importance of this gene in embryonic development. Additional isoforms resulting from alternatively spliced transcript variants, and use of upstream non-AUG (CUG) start codon have been reported for this gene. |
| Genomic regions | Chromosome 7 |
Strategy Summary
This gene has 7 protein coding transcripts:
| Name | Transcript ID | bp | Protein | Biotype | CCDS | UniProt Match | RefSeq Match | Flags |
| PEG10-202 | ENST00000482108.1 | 6618 | 325aa | Protein coding | CCDS55126 | Q86TG7-2 | - | TSL:1, GENCODE basic, APPRIS ALT2, |
| PEG10-208 | ENST00000615790.5 | 6394 | 359aa | Protein coding | CCDS75637 | A0A087WZG9 | - | TSL:1, GENCODE basic, APPRIS ALT2, |
| PEG10-203 | ENST00000488574.5 | 2587 | 401aa | Protein coding | CCDS75636 | B4DSP0 | - | TSL:2, GENCODE basic, APPRIS P4, |
| PEG10-206 | ENST00000612941.2 | 6392 | 741aa | Protein coding | - | A0A087WUL4 | - | TSL:5, GENCODE basic, APPRIS ALT2, |
| PEG10-209 | ENST00000617526.4 | 6392 | 707aa | Protein coding | - | A0A087WXK2 | - | TSL:5, GENCODE basic, APPRIS ALT2, |
| PEG10-205 | ENST00000612748.1 | 2585 | 783aa | Protein coding | - | A0A087WX23 | - | TSL:5, GENCODE basic, APPRIS ALT2, |
| PEG10-207 | ENST00000613043.1 | 574 | 31aa | Protein coding | - | A0A087WYS2 | - | CDS 3' incomplete, TSL:5, |
| PEG10-201 | ENST00000465184.1 | 788 | No protein | Processed transcript | - | - | - | TSL:3, |
| PEG10-204 | ENST00000493935.1 | 763 | No protein | Processed transcript | - | - | - | TSL:4, |




Red Cotton™ Assessment
Project Difficulty Level unknown
| Target Gene | PEG10 |
| This KO Strategy | loading |
| Red Cotton™ Notes | Gene PEG10 had been KO in hela cell line. |
Aforementioned information comes from Ubigene database. Different origin of cell lines may have
different condition. Ubigene reserved all the right for final explanation.
Special deals for this gene:
$49
Single gRNA plasmid off-shelf
$599
Single gRNA lentivirus



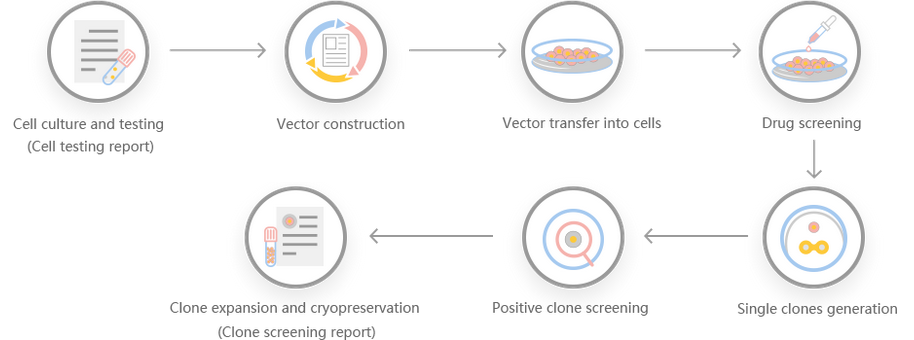
Work flow






 Comment:
Comment: