CRISPR-U™ Gene Knockout Cell Line
Strategy
FGFR1 Gene Knockout Strategy
CRISPR-U™ technology (CRISPR based), developed by Ubigene, is more efficient than general
CRISPR/Cas9 technology in double-strand breaking and homologous recombination. With CRISPR-U™, Ubigene has
successfully edited over 3000 genes on more than 100 types of cell lines.
Objective
To create a Human FGFR1 Knockout
model in cell line by
CRISPR-U™-mediated
genome
engineering.
Target gene info 
| Official symbol | FGFR1 |
| Gene id | 2260 |
| Organism | Homo sapiens |
| Official full symbol | fibroblast growth factor receptor 1 |
| Gene type | protein-coding |
| Also known as | BFGFR, CD331, CEK, ECCL, FGFBR, FGFR-1, FLG, FLT-2, FLT2, HBGFR, HH2, HRTFDS, KAL2, N-SAM, OGD, bFGF-R-1 |
| Summary | The protein encoded by this gene is a member of the fibroblast growth factor receptor (FGFR) family, where amino acid sequence is highly conserved between members and throughout evolution. FGFR family members differ from one another in their ligand affinities and tissue distribution. A full-length representative protein consists of an extracellular region, composed of three immunoglobulin-like domains, a single hydrophobic membrane-spanning segment and a cytoplasmic tyrosine kinase domain. The extracellular portion of the protein interacts with fibroblast growth factors, setting in motion a cascade of downstream signals, ultimately influencing mitogenesis and differentiation. This particular family member binds both acidic and basic fibroblast growth factors and is involved in limb induction. Mutations in this gene have been associated with Pfeiffer syndrome, Jackson-Weiss syndrome, Antley-Bixler syndrome, osteoglophonic dysplasia, and autosomal dominant Kallmann syndrome 2. Chromosomal aberrations involving this gene are associated with stem cell myeloproliferative disorder and stem cell leukemia lymphoma syndrome. Alternatively spliced variants which encode different protein isoforms have been described; however, not all variants have been fully characterized. |
| Genomic regions | Chromosome 8 |
Strategy Summary
This gene has 20 protein coding transcripts:
| Name | Transcript ID | bp | Protein | Biotype | CCDS | UniProt Match | RefSeq Match | Flags |
| FGFR1-206 | ENST00000397091.9 | 5702 | 820aa | Protein coding | CCDS43732 | P11362-7 | - | TSL:1, GENCODE basic, APPRIS ALT1, |
| FGFR1-214 | ENST00000447712.7 | 5697 | 822aa | Protein coding | CCDS6107 | P11362-1 | NM_023110.3 | TSL:1, GENCODE basic, APPRIS ALT1, MANE Select v0.92, |
| FGFR1-238 | ENST00000532791.5 | 5590 | 820aa | Protein coding | CCDS55222 | P11362-2 | - | TSL:5, GENCODE basic, APPRIS P4, |
| FGFR1-211 | ENST00000425967.7 | 5375 | 853aa | Protein coding | CCDS55223 | P11362-21 | - | TSL:1, GENCODE basic, |
| FGFR1-202 | ENST00000335922.9 | 4157 | 812aa | Protein coding | CCDS55221 | P11362-20 | - | TSL:1, GENCODE basic, |
| FGFR1-204 | ENST00000356207.9 | 3824 | 733aa | Protein coding | CCDS43730 | P11362-3 | - | TSL:1, GENCODE basic, |
| FGFR1-201 | ENST00000326324.10 | 3816 | 731aa | Protein coding | CCDS43731 | P11362-14 | - | TSL:1, GENCODE basic, |
| FGFR1-209 | ENST00000397113.6 | 3680 | 820aa | Protein coding | CCDS43732 | P11362-7 | - | TSL:2, GENCODE basic, APPRIS ALT1, |
| FGFR1-208 | ENST00000397108.8 | 2870 | 820aa | Protein coding | CCDS43732 | P11362-7 | - | TSL:1, GENCODE basic, APPRIS ALT1, |
| FGFR1-203 | ENST00000341462.8 | 6054 | 367aa | Protein coding | - | C1KBH7 | - | TSL:5, GENCODE basic, |
| FGFR1-242 | ENST00000619564.3 | 5482 | 228aa | Protein coding | - | B5A958 | - | TSL:5, GENCODE basic, |
| FGFR1-207 | ENST00000397103.5 | 2953 | 733aa | Protein coding | - | E7EU09 | - | TSL:5, GENCODE basic, |
| FGFR1-226 | ENST00000525001.5 | 1155 | 298aa | Protein coding | - | E9PNM3 | - | CDS 3' incomplete, TSL:5, |
| FGFR1-233 | ENST00000529552.5 | 688 | 142aa | Protein coding | - | E9PKV7 | - | CDS 3' incomplete, TSL:4, |
| FGFR1-229 | ENST00000526742.5 | 657 | 145aa | Protein coding | - | E9PKF2 | - | CDS 3' incomplete, TSL:4, |
| FGFR1-210 | ENST00000413133.6 | 587 | 106aa | Protein coding | - | C9J205 | - | CDS 3' incomplete, TSL:4, |
| FGFR1-241 | ENST00000533668.5 | 571 | 139aa | Protein coding | - | E9PN14 | - | CDS 3' incomplete, TSL:5, |
| FGFR1-212 | ENST00000434187.5 | 529 | 54aa | Protein coding | - | C9J1L5 | - | CDS 3' incomplete, TSL:4, |
| FGFR1-234 | ENST00000530568.5 | 467 | 85aa | Protein coding | - | E9PQ40 | - | CDS 3' incomplete, TSL:3, |
| FGFR1-213 | ENST00000440174.1 | 389 | 1aa | Protein coding | - | - | - | CDS 3' incomplete, TSL:3, |
| FGFR1-243 | ENST00000649678.1 | 6216 | 818aa | Nonsense mediated decay | - | A0A3B3ISD1 | - | - |
| FGFR1-247 | ENST00000674380.1 | 5002 | 42aa | Nonsense mediated decay | - | A0A6I8PTV4 | - | - |
| FGFR1-244 | ENST00000674189.1 | 3309 | 42aa | Nonsense mediated decay | - | A0A6I8PTV4 | - | - |
| FGFR1-222 | ENST00000487647.5 | 1608 | 61aa | Nonsense mediated decay | - | E9PKX3 | - | TSL:1, |
| FGFR1-246 | ENST00000674235.1 | 1071 | 245aa | Nonsense mediated decay | - | A0A6I8PRY1 | - | CDS 5' incomplete, |
| FGFR1-236 | ENST00000531196.5 | 842 | 134aa | Nonsense mediated decay | - | H0YE20 | - | CDS 5' incomplete, TSL:3, |
| FGFR1-221 | ENST00000484370.5 | 826 | 150aa | Nonsense mediated decay | - | P11362-15 | - | TSL:1, |
| FGFR1-224 | ENST00000496629.1 | 584 | No protein | Processed transcript | - | - | - | TSL:4, |
| FGFR1-228 | ENST00000526688.1 | 570 | No protein | Processed transcript | - | - | - | TSL:3, |
| FGFR1-231 | ENST00000527203.5 | 556 | No protein | Processed transcript | - | - | - | TSL:3, |
| FGFR1-220 | ENST00000480571.1 | 442 | No protein | Processed transcript | - | - | - | TSL:4, |
| FGFR1-239 | ENST00000533301.1 | 377 | No protein | Processed transcript | - | - | - | TSL:3, |
| FGFR1-235 | ENST00000530701.1 | 276 | No protein | Processed transcript | - | - | - | TSL:1, |
| FGFR1-227 | ENST00000526570.5 | 5528 | No protein | Retained intron | - | - | - | TSL:1, |
| FGFR1-248 | ENST00000674474.1 | 4711 | No protein | Retained intron | - | - | - | - |
| FGFR1-217 | ENST00000470826.5 | 2281 | No protein | Retained intron | - | - | - | TSL:1, |
| FGFR1-245 | ENST00000674217.1 | 2011 | No protein | Retained intron | - | - | - | - |
| FGFR1-223 | ENST00000496296.5 | 1898 | No protein | Retained intron | - | - | - | TSL:1, |
| FGFR1-230 | ENST00000527114.5 | 1499 | No protein | Retained intron | - | - | - | TSL:5, |
| FGFR1-216 | ENST00000466021.5 | 823 | No protein | Retained intron | - | - | - | TSL:2, |
| FGFR1-232 | ENST00000527745.3 | 790 | No protein | Retained intron | - | - | - | TSL:3, |
| FGFR1-205 | ENST00000397090.4 | 777 | No protein | Retained intron | - | - | - | TSL:2, |
| FGFR1-218 | ENST00000474970.1 | 764 | No protein | Retained intron | - | - | - | TSL:5, |
| FGFR1-237 | ENST00000532386.5 | 725 | No protein | Retained intron | - | - | - | TSL:3, |
| FGFR1-215 | ENST00000464163.1 | 545 | No protein | Retained intron | - | - | - | TSL:4, |
| FGFR1-240 | ENST00000533619.5 | 540 | No protein | Retained intron | - | - | - | TSL:4, |
| FGFR1-225 | ENST00000524528.1 | 486 | No protein | Retained intron | - | - | - | TSL:3, |
| FGFR1-219 | ENST00000475621.1 | 485 | No protein | Retained intron | - | - | - | TSL:2, |




Red Cotton™ Assessment
Project Difficulty Level unknown
| Target Gene | FGFR1 |
| This KO Strategy | loading |
| Red Cotton™ Notes | Gene FGFR1 had been KO in hek293t cell line. |
Aforementioned information comes from Ubigene database. Different origin of cell lines may have
different condition. Ubigene reserved all the right for final explanation.
Special deals for this gene:
$49
Single gRNA plasmid off-shelf
$599
Single gRNA lentivirus



Work flow
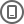
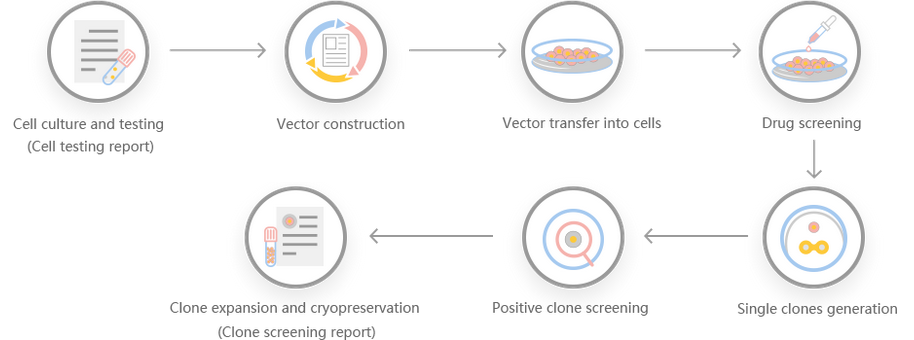






 Comment:
Comment: